Rethinking LLM Evaluation: Can We Evaluate LLMs with 200× Less Data?
Introducing EssenceBench, a coarse-to-fine framework for LLM benchmark compression using iterative Genetic Algorithms.
Rethinking LLM Evaluation: Can We Evaluate LLMs with 200× Less Data?
Authors: Shaobo Wang*, Cong Wang*, Wenjie Fu*, Yue Min, Mingquan Feng, Isabel Guan, Xuming Hu, Conghui He, Cunxiang Wang, Kexin Yang, Xingzhang Ren, Fei Huang, Dayiheng Liu, Linfeng Zhang†
Affiliations: Shanghai Jiao Tong University, Alibaba Qwen Team, Fudan University, Harbin Institute of Technology, Hong Kong University of Science and Technology, Hong Kong University of Science and Technology(Guangzhou), Zhipu AI, Shanghai AI Lab
As the demand for comprehensive evaluations of diverse model capabilities steadily increases, benchmark suites have correspondingly grown significantly in scale. Despite notable advances in redundancy reduction, a systematic framework that ensures both prediction accuracy and ranking consistency is still largely elusive. In this paper, we propose EssenceBench, a coarse-to-fine framework utilizing an iterative Genetic Algorithm (GA), which takes the advantages of fitness-based subset search and attribution-based sample search.
Our approach yields superior compression results with lower reconstruction error and markedly higher efficiency. In particular, on the HellaSwag benchmark (10K samples), our method preserves the ranking of all models shifting within 5% using 25× fewer samples, and achieves 95% ranking preservation shifting within 5% using only 200× fewer samples.
Overview
In recent years, LLM benchmarks have shifted from traditional NLP tasks to more comprehensive, multidimensional evaluation suites. However, as the scope and granularity of evaluation expand, so does the scale and computing cost. For instance, evaluating a model like Qwen2.5-7B-Instruct across over 60 subtasks in OpenCompass often takes about 1k GPU hours.
We revisit the foundations of LLM benchmark evaluation, focusing on a critical yet under-examined phenomenon: sample redundancy. Our analysis reveals that many benchmarks exhibit high overlap both in the text of their prompts and in the models’ performance rankings. To address this, we frame benchmark compression as an optimization problem with the aim of score reconstruction.
1. Sample Redundancy
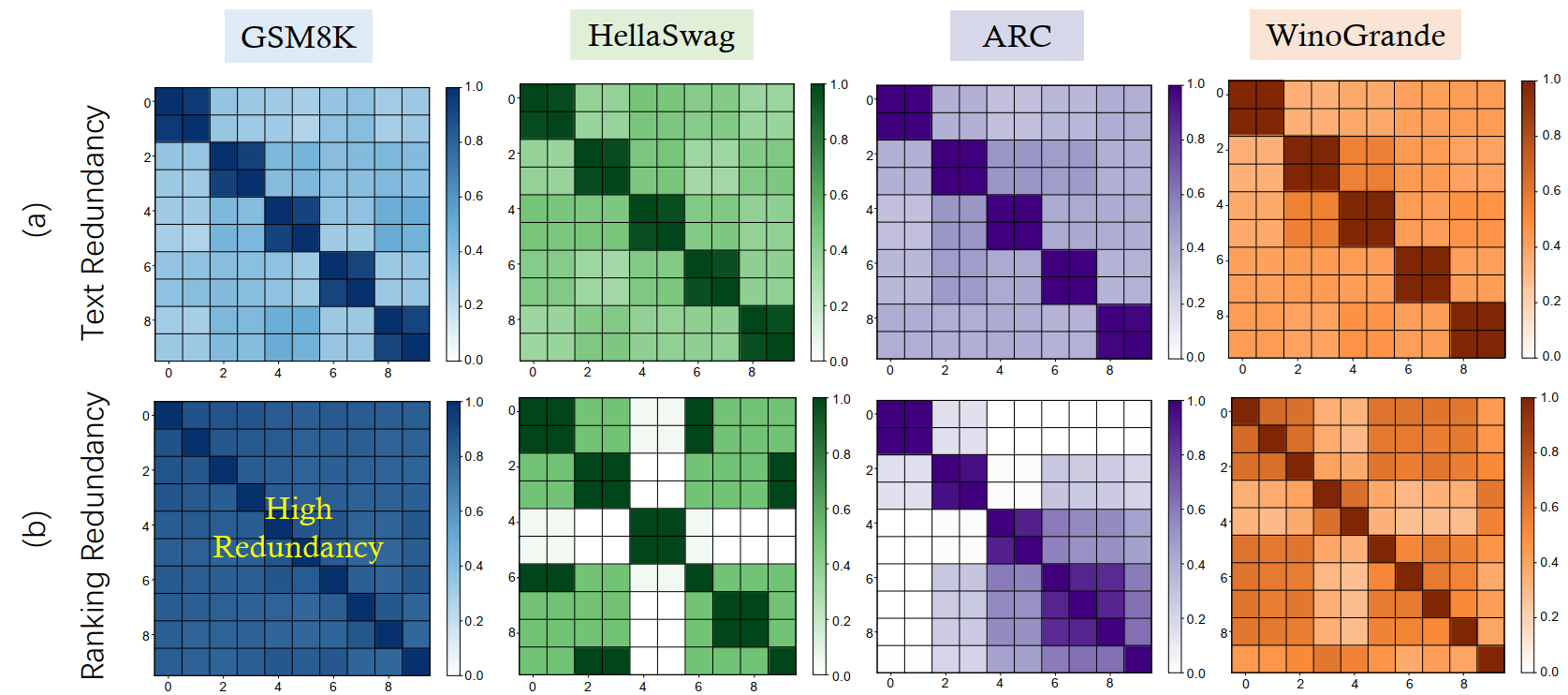
Intuitively, when two examples share similar wording or contain nearly identical behavior across models, retaining both adds little new information while doubling evaluation costs. We quantify this through two dimensions:
1.1 Text-level Redundancy
Defined as lexical and semantic overlap between evaluation instances. For a sample pair $(x_i, x_j)$, the redundancy is: \(\small \mathcal{R}_{\text{text}}(i, j) = \langle \text{Emb}(x_i), \text{Emb}(x_j) \rangle\) where $\text{Emb}(\cdot)$ denotes an embedding mapping.
1.2 Ranking-level Redundancy
Measured through the consistency of model performance rankings across samples. The redundancy between two samples $x_i$ and $x_j$ is quantified by the Pearson correlation coefficient of their performance vectors $s_i, s_j$: \(\small \mathcal{R}_{\text{ranking}}(i, j) = \rho(s_i, s_j) = \frac{\text{Cov}(s_i, s_j)}{\sigma_{s_i}\sigma_{s_j}}\) A high correlation indicates that the set of models exhibits an identical performance pattern on both samples, implying they provide redundant information regarding model ranking capabilities.
2. EssenceBench
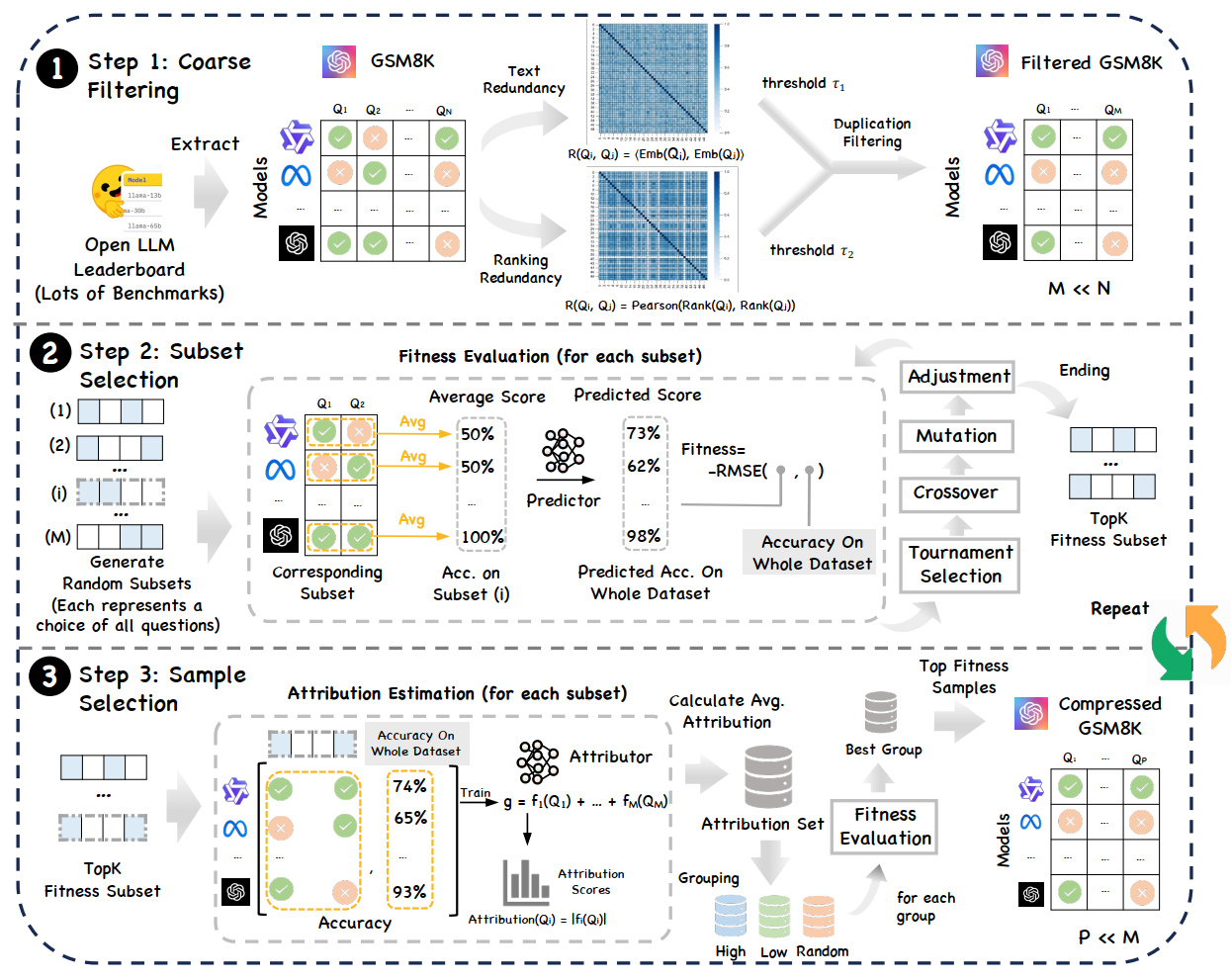
The benchmark compression problem is framed as selecting a $k$-element subset $\tilde{\mathcal{D}}$ that best reconstructs the full-dataset score: \(\small \tilde{\mathcal{D}}^* = \arg\min_{\tilde{\mathcal{D}} \subseteq \mathcal{D}, |\tilde{\mathcal{D}}|=k} \mathcal{L}(g(\mathcal{D}), g(\tilde{\mathcal{D}}))\)
2.1 Step 1. Coarse Filtering
Based on text and ranking redundancy thresholds ($\tau_{\text{text}}$, $\tau_{\text{ranking}}$), we examine the dataset in its original order. If a sample $x_i$ is highly redundant with any previously kept sample, it is discarded. This strips away the most redundant examples to yield a compact yet representative filtered set of size $M$.
2.2 Step 2. Fitness-based Subset Selection
We employ an iterative Genetic Algorithm (GA) to search for high-quality subsets.
- Fitness Evaluation: We train a General Additive Model (GAM) once at the beginning of each GA round to act as a surrogate scoring function $g$.
- Evolution: Starting from random $k$-element masks, we perform tournament selection, crossover, and mutation to minimize the RMSE between predicted subset scores and ground-truth full-set scores.
2.3 Step 3. Attribution-based Sample Selection
To maximize reconstruction fidelity, we train an Explainable Boosting Machine (EBM) on the elite masks from Step 2. Each sample is assigned an attribution score $A_l$: \(\small A_l = \frac{\sum_{m \in \mathcal{E}} 1\{l \in \mathcal{I}(m)\} \|f_l^m\|_2}{\sum_{m \in \mathcal{E}} 1\{l \in \mathcal{I}(m)\}}\) Samples are stratified into “High”, “Low”, and “Random” attribution groups. GA is then reapplied within these groups to identify the most representative and informative subset, ensuring sample-level diversity and alleviating suboptimal convergence.
3. Experimental Results
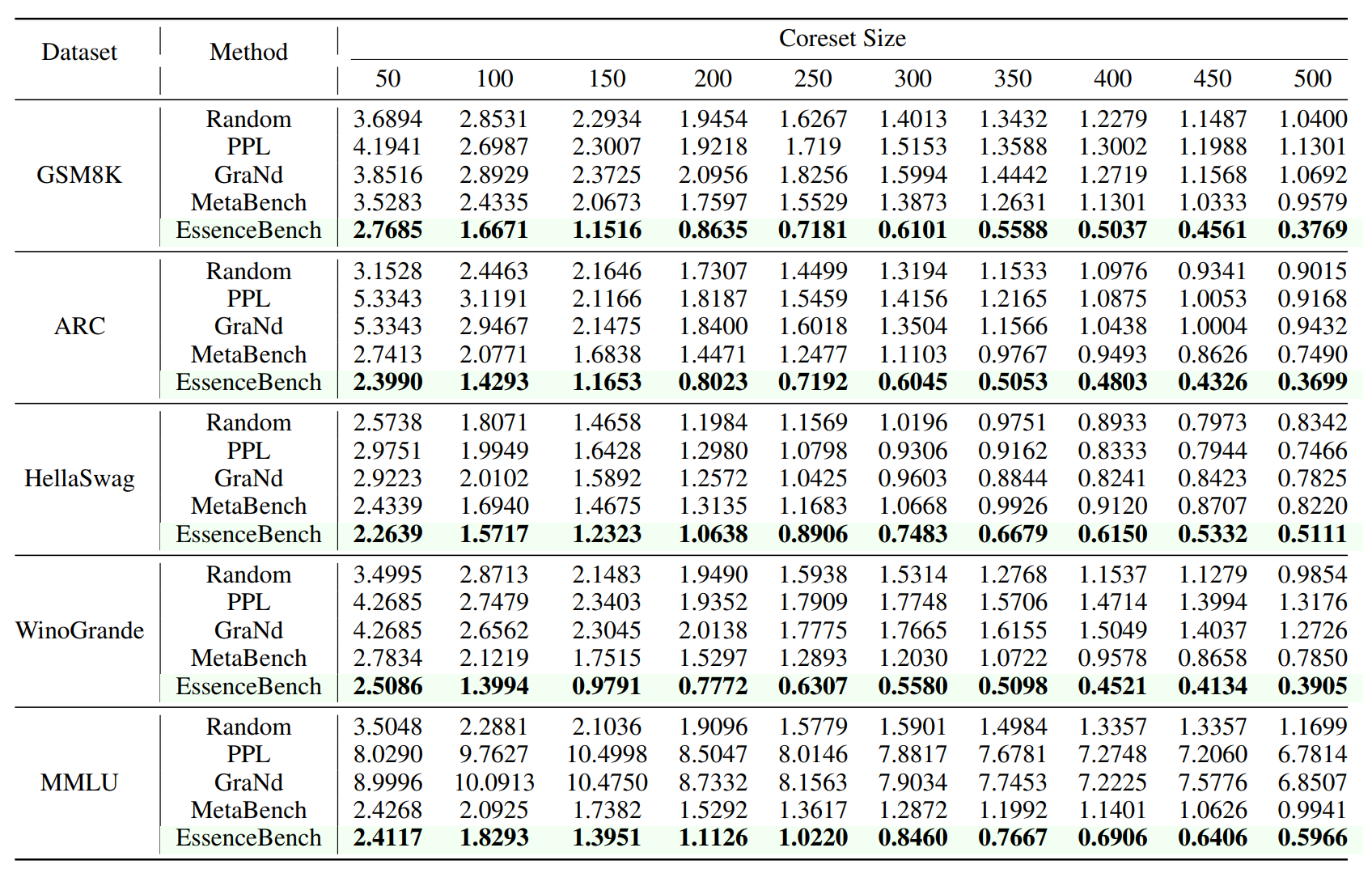
3.1 Better Performance with Same Compression
On five major benchmarks (GSM8K, ARC, HellaSwag, WinoGrande, MMLU), EssenceBench achieves state-of-the-art results. Notably, on GSM8K with a subset size of 500, our method achieves a 60.7% reduction in RMSE compared to MetaBench.
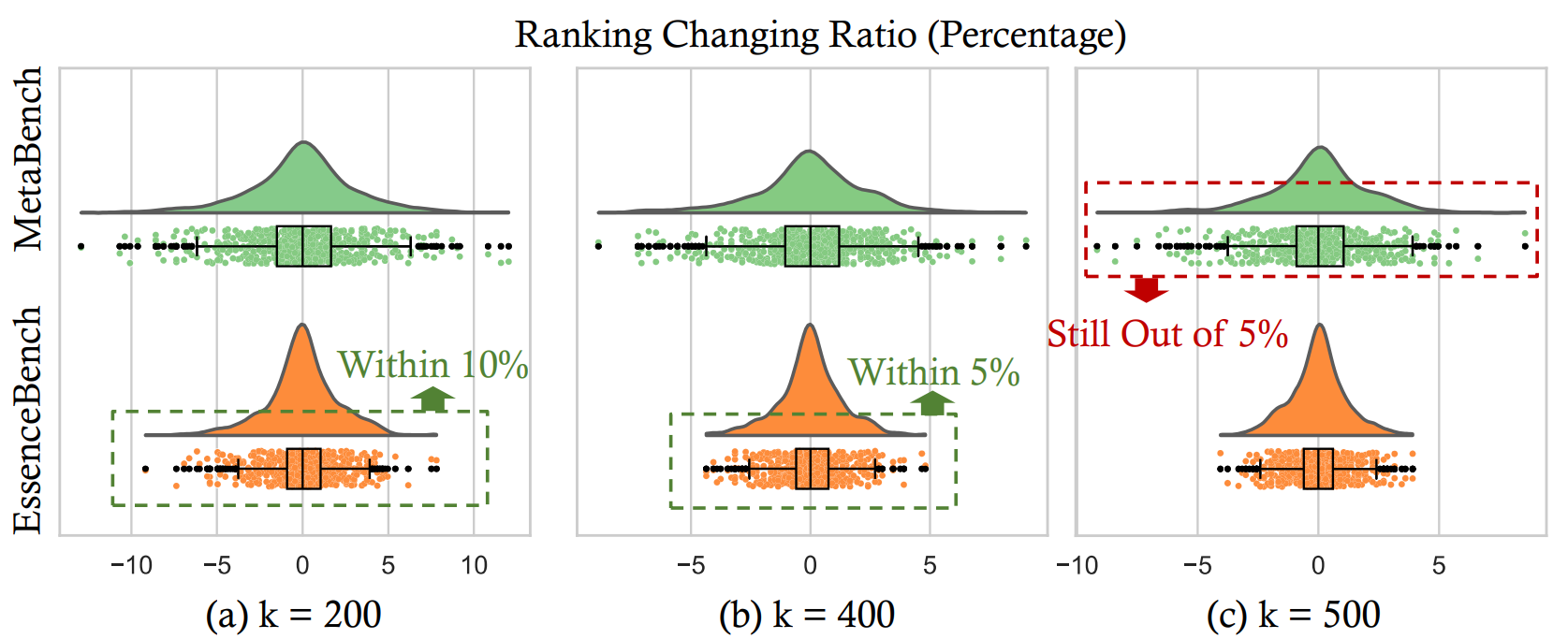
3.2 Ranking Fidelity at 200× Reduction
Notably, we found that selecting just 200× less samples (e.g., $k=50$ for HellaSwag) preserves 95% of rankings changing within 10%. With $k=400$, all ranking shifts remain within 5%, offering significantly tighter preservation than previous approaches.
4. Ablation Studies and Efficiency
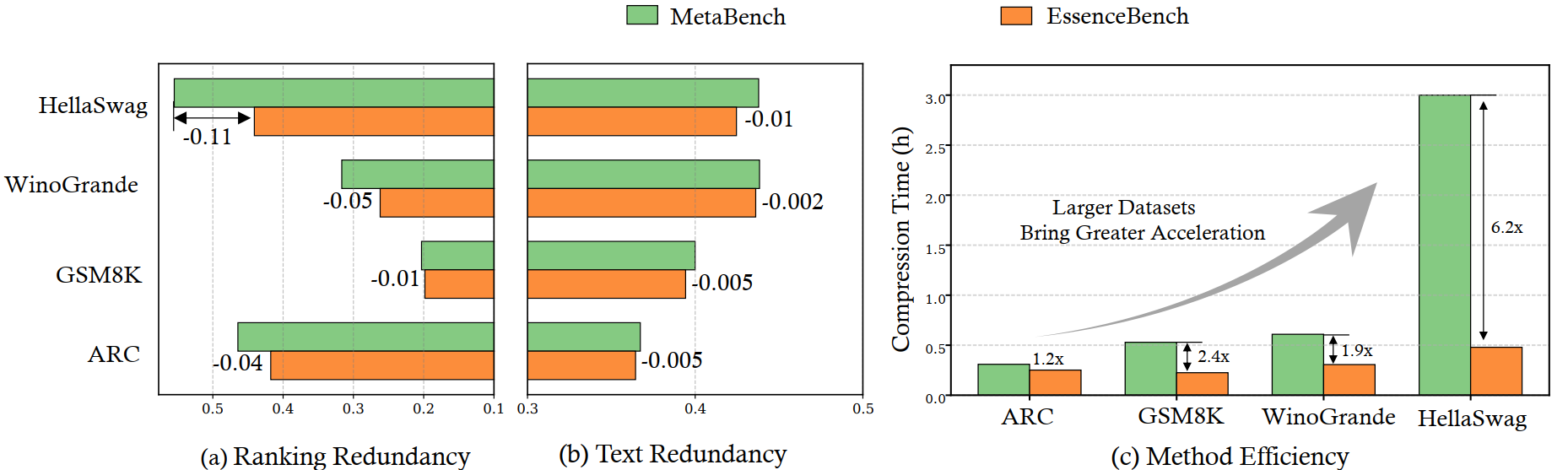
4.1 Computational Efficiency
The speedup ratio of EssenceBench over MetaBench increases as the dataset size grows. On HellaSwag, EssenceBench is 6.2× faster than MetaBench, primarily due to the coarse filtering stage and the use of the GAM surrogate which avoids per-candidate retraining.
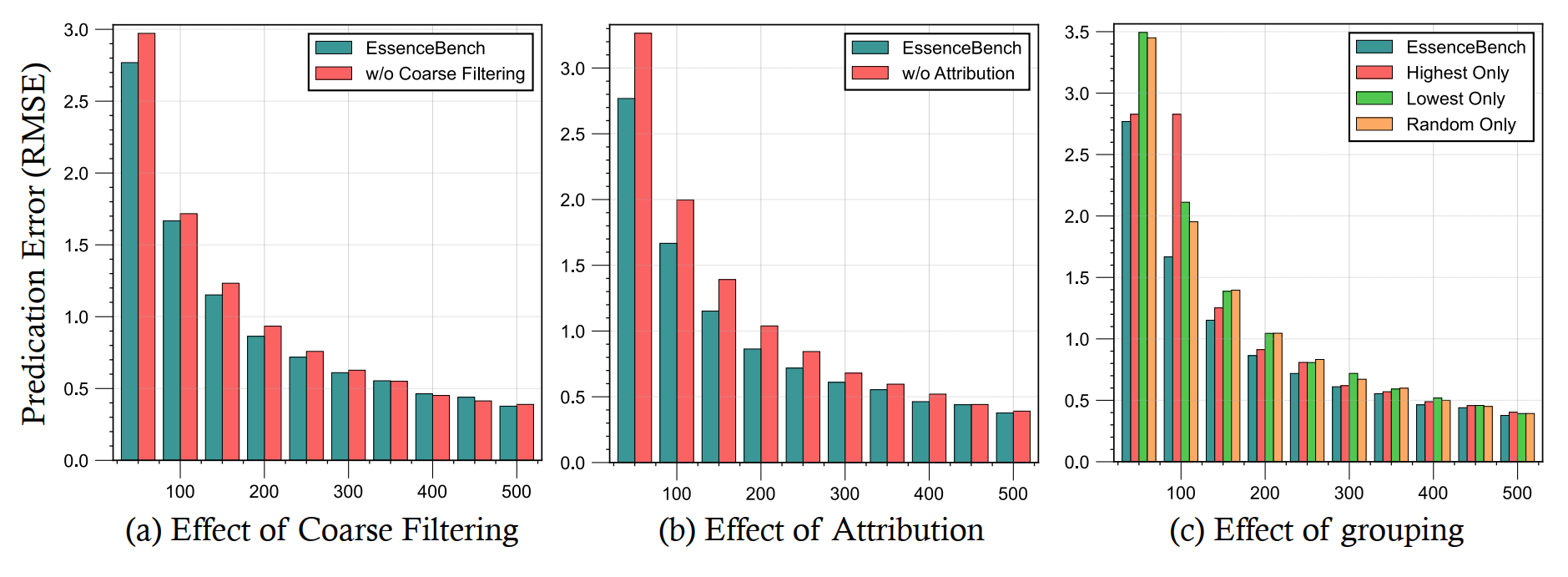
4.2 Impact of Attribution and Grouping
Our ablation studies on GSM8K show:
- Coarse Filtering: significantly reduces RMSE, especially when the subset size is small.
- Attribution: outperforms selection without attribution when the subset size is below 400.
- Grouping: Effectively enhances compression quality by ensuring fine-grained diversity.
5. Conclusion
In this paper, we identified sample redundancy in LLM benchmark evaluation. To address this problem, we introduced EssenceBench, a coarse-to-fine benchmark compression framework that combines redundancy-aware filtering with an iterative genetic algorithm optimized for accurate reconstruction. Extensive experiments on five standard benchmarks show that EssenceBench achieves a 200× reduction in benchmark size while preserving ranking fidelity.
Citation
@article{wang2025rethinking,
title={Rethinking LLM Evaluation: Can We Evaluate LLMs with 200x Less Data?},
author={Wang, Shaobo and Wang, Cong and Fu, Wenjie and Min, Yue and Feng, Mingquan and Guan, Isabel and Hu, Xuming and He, Conghui and Wang, Cunxiang and Yang, Kexin and others},
journal={arXiv preprint arXiv:2510.10457},
year={2025}
}